Goal
This vignette introduces the mashr software for someone
who is familiar with the basic idea in Urbut et
al., which we strongly recommend reading before proceeding.
In this first vignette we use only the “canonical” covariance matries
to simplify presentation. However, an important aspect of
mashr is the use of data-driven covariance matrices that
are learned from the data, so when you are done here you should look at
the data-driven vignette and the non-canonical matrix vignette. If you
think you may have correlations in your data measurements then also look
at how to incorporate correlations.
Outline
This package implements the “multivariate adaptive shrinkage” (mash) method from Urbut et al..
Mash is a method for dealing with large numbers of tests in many (e.g. dozens) different conditions. Here “conditions” can mean many different things: it could be different tissues (as in Urbut et al), or different time points, or different treatments, or even different phenotypes (think of testing multiple phenotypes at many genetic variants in a GWAS).
There are essentially four steps to a mash analysis
- Read in the data
- Set up the covariance matrices to be used
- Fit the model
- Extract posterior summaries and other quantities of interest
Here we go through an example that illustrates each of these steps in turn. In this example we use the same (simulated) data at each step. In complex practical applications – especially very large applications – one might want to use different subsets of the data at different steps. For example, one might only want to extract posterior summaries (step 4) for a subset of the tests. We will deal with such complications in later vignettes. For now, we note the one crucial rule:
The Crucial Rule: Step 3 (fitting the mash
model) must be performed with either all the tests you
performed, or – if that causes computational challenges – a large
random subset of tests.
In particular, you must not use only a set of selected “significant”
or “strong” tests in step 3. This is because mash uses step
3 to learn about the null signal in the data, as well as the non-null
signal. In particular, in settings where most tests are null or
nearly-null, this is the point where mash learns this, and
consequently “shrinks” (“corrects”) the posterior estimates towards 0.
(Effectively this step is analogous to a “multiple testing correction”
step.)
Simulate some data
First we simulate some data for illustration.
This simulation routine creates a dataset with 5 conditions, and four different types of effect: null, independent among conditions, condition-specific in condition 1, and shared (equal effects in all conditions). It creates 500 effects of each type for a total of 2000 effects.
Step 1: Read in the data
To run mash you need data consisting of a matrix of
effects (Bhat) and a matrix of standard errors
(Shat), for \(J\) effects
(rows) in \(R\) conditions
(columns).
[If you have only access to \(Z\)
scores, you can set Bhat to the Z scores, and set
Shat to be the matrix with all 1s].
The simulation above created both these matrices for us (in
simdata$Bhat and simdata$Shat). To get these
ready for applying mash you must first use
mash_set_data to create a data object with those two pieces
of information:
data = mash_set_data(simdata$Bhat, simdata$Shat)Step 2: Set up the covariance matrices
There are two types of covariance matrix you can use in
mash: “canonical” and “data-driven”. The canonical ones are
very easy to set up and so we use those here for illustration. However,
in applications you will likely also want to use data-driven matrices,
and this is an important feature of mash. See the data-driven vignette for more details
on how to do this.
The function to set up canonical covariance matries is
cov_canonical. The following sets up canonical covariances
in U.c (we used .c to indicate canonical),
which is a named list of matrices.
U.c = cov_canonical(data)
print(names(U.c))# [1] "identity" "condition_1" "condition_2" "condition_3"
# [5] "condition_4" "condition_5" "equal_effects" "simple_het_1"
# [9] "simple_het_2" "simple_het_3"Step 3: fit the model
Having set up the data and covariance matrices you are ready to fit
the model using the mash function:
m.c = mash(data, U.c)# - Computing 2000 x 151 likelihood matrix.
# - Likelihood calculations took 0.05 seconds.
# - Fitting model with 151 mixture components.
# - Model fitting took 0.54 seconds.
# - Computing posterior matrices.
# - Computation allocated took 0.02 seconds.This can take a little time. What this does is to fit a mixture model
to the data, estimating the mixture proportions. Specifically the model
is that the true effects follow a mixture of multivariate normal
distributions: \(B \sim \sum_k \sum_l \pi_{kl}
N(0, \omega_l U_k)\) where the \(\omega_l\) are scaling factors set by the
“grid” parameter in mash and the \(U_k\) are the covariance matrices (here
specified by U.c).
Remember the Crucial Rule! This step must be peformed using all the
tests (or a large random subset), because this is where
mash learns that many tests are null and corrects for
it.
Step 4: Extract Posterior Summaries
You can extract estimates (posterior means and posterior standard
deviations) and measures of significance (local false sign rates) using
functions like get_pm (posterior mean),
get_psd (posteriore standard deviation) and
get_lfsr (local false sign rate):
# condition_1 condition_2 condition_3 condition_4 condition_5
# effect_1 0.7561458 0.7705833 0.8052784 0.8111106 0.8189440
# effect_2 0.7245445 0.6905871 0.7760428 0.6922674 0.7012653
# effect_3 0.7387335 0.7514386 0.8104767 0.8390446 0.8501584
# effect_4 0.7855682 0.8454469 0.8360452 0.8660075 0.8503394
# effect_5 0.8044530 0.8371033 0.8808419 0.8785890 0.8758291
# effect_6 0.7206973 0.6833578 0.7894008 0.7002639 0.7569357# condition_1 condition_2 condition_3 condition_4 condition_5
# effect_1 0.07387924 -0.12489892 -0.0763266677 0.092636122 -0.045368333
# effect_2 -0.02353728 -0.17604903 -0.0324348873 -0.201656883 -0.170553647
# effect_3 -0.10218378 0.18092246 -0.0846361802 0.022525113 -0.002684966
# effect_4 -0.06238618 0.02794225 0.0654499922 0.008113534 0.032179633
# effect_5 0.03341406 -0.06059792 0.0001472222 -0.003887722 -0.004665907
# effect_6 -0.09934823 0.21556824 -0.0791530347 0.213031210 0.079063543# condition_1 condition_2 condition_3 condition_4 condition_5
# effect_1 0.4592858 0.4210499 0.3544674 0.4062428 0.3365922
# effect_2 0.4861870 0.4277966 0.4047004 0.4447897 0.4149876
# effect_3 0.4679453 0.5082149 0.3799464 0.3269399 0.3283911
# effect_4 0.4389151 0.2964005 0.3082095 0.2749191 0.2809670
# effect_5 0.4033380 0.3286542 0.2608065 0.2597960 0.2656089
# effect_6 0.5226691 0.4981609 0.4482801 0.4906789 0.3899259Each of these are \(J \times R\) matrices.
Use get_significant_results to find the indices of
effects that are “significant”, which here means they have lfsr less
than t in at least one condition, where t is a threshold you specify
(default 0.05). The output is ordered from most significant to least
significant.
head(get_significant_results(m.c))# effect_1800 effect_1812 effect_1857 effect_1953 effect_1915 effect_1950
# 1800 1812 1857 1953 1915 1950
print(length(get_significant_results(m.c)))# [1] 150You can also get the significant results in just a subset of conditions. For example
print(head(get_significant_results(m.c, conditions=1)))# effect_1812 effect_1726 effect_1800 effect_1501 effect_1953 effect_1993
# 1812 1726 1800 1501 1953 1993Sharing
Use get_pairwise_sharing to assess sharing of
significant signals among each pair of conditions. Here the default
definition of shared is “the same sign and within a factor 0.5 of each
other”.
print(get_pairwise_sharing(m.c)) # condition_1 condition_2 condition_3 condition_4 condition_5
# condition_1 1.0000000 0.7761194 0.8015267 0.7727273 0.7777778
# condition_2 0.7761194 1.0000000 0.9142857 0.8971963 0.8611111
# condition_3 0.8015267 0.9142857 1.0000000 0.9108911 0.8962264
# condition_4 0.7727273 0.8971963 0.9108911 1.0000000 0.8679245
# condition_5 0.7777778 0.8611111 0.8962264 0.8679245 1.0000000You can change the factor if you like. For example, here by setting the factor to be 0 you assess only if they are the same sign:
print(get_pairwise_sharing(m.c, factor=0))# condition_1 condition_2 condition_3 condition_4 condition_5
# condition_1 1.0000000 0.8880597 0.9236641 0.8863636 0.9111111
# condition_2 0.8880597 1.0000000 0.9333333 0.9252336 0.9444444
# condition_3 0.9236641 0.9333333 1.0000000 0.9207921 0.9716981
# condition_4 0.8863636 0.9252336 0.9207921 1.0000000 0.9339623
# condition_5 0.9111111 0.9444444 0.9716981 0.9339623 1.0000000Measure of fit (log-likelihood)
Use get_loglik to find the log-likelihood of the fit
(this will only be useful when you have other fits to compare it
with!)
print(get_loglik(m.c))# [1] -16120.32Estimated mixture proportions
Use get_estimated_pi to extract the estimates of the
mixture proportions for different types of covariance matrix:
print(get_estimated_pi(m.c))# null identity condition_1 condition_2 condition_3
# 3.711961e-01 2.208687e-01 1.365259e-01 2.097879e-02 0.000000e+00
# condition_4 condition_5 equal_effects simple_het_1 simple_het_2
# 0.000000e+00 4.959624e-03 2.264402e-01 8.758416e-05 0.000000e+00
# simple_het_3
# 1.894312e-02
barplot(get_estimated_pi(m.c),las = 2)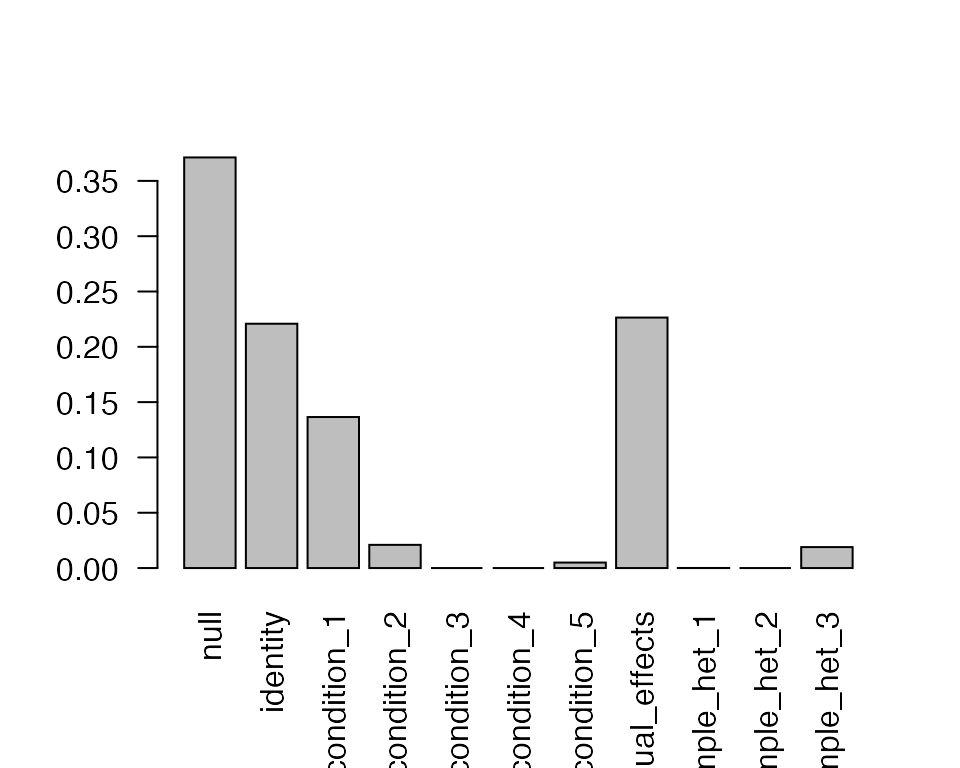
Here we can see most of the mass is on the null, identity,
singletons_1 (which corresponds to effects that are
specific to condition 1) and equal_effects. This
reassuringly matches the way that these data were generated.
Metaplot
The following produces a meta-plot based on the posterior means and posterior variances of an effect. Here we look at the most significant result.
mash_plot_meta(m.c,get_significant_results(m.c)[1])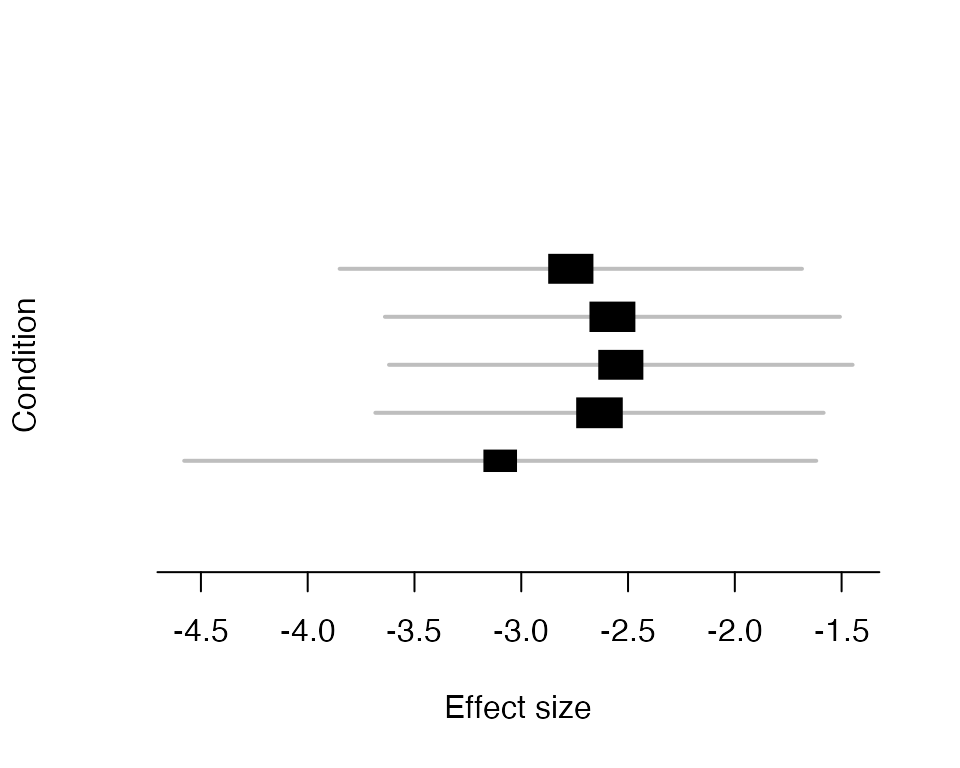
Session information.
print(sessionInfo())# R version 3.6.2 (2019-12-12)
# Platform: x86_64-apple-darwin15.6.0 (64-bit)
# Running under: macOS Catalina 10.15.7
#
# Matrix products: default
# BLAS: /Library/Frameworks/R.framework/Versions/3.6/Resources/lib/libRblas.0.dylib
# LAPACK: /Library/Frameworks/R.framework/Versions/3.6/Resources/lib/libRlapack.dylib
#
# locale:
# [1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
#
# attached base packages:
# [1] stats graphics grDevices utils datasets methods base
#
# other attached packages:
# [1] mashr_0.2.73 ashr_2.2-57
#
# loaded via a namespace (and not attached):
# [1] Rcpp_1.0.8 highr_0.8 bslib_0.3.1 compiler_3.6.2
# [5] jquerylib_0.1.4 plyr_1.8.5 tools_3.6.2 digest_0.6.23
# [9] jsonlite_1.7.2 evaluate_0.14 memoise_1.1.0 lattice_0.20-38
# [13] rlang_1.0.6 Matrix_1.3-4 cli_3.5.0 yaml_2.2.0
# [17] mvtnorm_1.0-11 pkgdown_2.0.7 xfun_0.36 fastmap_1.1.0
# [21] invgamma_1.1 stringr_1.4.0 knitr_1.37 desc_1.2.0
# [25] fs_1.5.2 sass_0.4.0 systemfonts_1.0.2 rprojroot_2.0.3
# [29] grid_3.6.2 R6_2.4.1 rmarkdown_2.21 mixsqp_0.3-48
# [33] rmeta_3.0 irlba_2.3.3 purrr_0.3.4 magrittr_2.0.1
# [37] htmltools_0.5.4 assertthat_0.2.1 abind_1.4-5 ragg_0.3.1
# [41] stringi_1.4.3 truncnorm_1.0-8 SQUAREM_2017.10-1 crayon_1.4.1